H3Africa Collaborative Research Center
Genomic Characterization and Surveillance of Microbial Threats in West Africa
The Goal: to develop innovative genomic tools and strategies that will reduce the devastating impact of viral diseases, like Ebola and Lassa fever, on West African populations.
Project Leads
The Problem
Fevers of unknown origins are incredibly difficult to diagnose, due to their vast diversity and complexity. As a result, recent viral outbreaks have had a devastating impact on African populations, highlighting vulnerabilities in African public health systems. Tools to enable the rapid, accurate diagnosis and analysis of patient samples were often not readily available in the areas affected by outbreaks, making it more difficult to reduce morbidity and mortality during recent epidemics.
Project Strategy
- Utilize modern genomic technologies to better understand the nature and extent of fevers of unknown origins, including viral infections in West Africa.
- Develop robust and cost-effective field diagnostics to swiftly and accurately capture the prevalence of these viruses in the population.
- Use genomic and immunological data to characterize the pathogen-host interaction and enhance clinical care.
Outcomes to Date
Viral Research
- Established modern genomics hubs in Nigeria, Senegal, and Sierra Leone that have the ability to identify and analyze unknown circulating viruses.
- Used these hubs to rapidly characterize and track emergent viruses and outbreaks in West Africa including Lassa Virus (Nigeria, Sierra Leone), Yellow Fever (Nigeria), Dengue Virus (Sengal). This brisk action enabled local governments to implement containment strategies and enhanced patient care to effectively reduce the public health impact.
- Helped contain and characterize the Ebola outbreak in Nigeria.
Technology Development
- Developed a novel variation of SHERLOCK, a low-cost genomics test, to rapidly identify three types of fever at the bedside, reducing the standard of care diagnostic time; deployed during the recent Lassa outbreak in Nigeria.
- Developed additional genomics tests to enable rapid identification of Ebola virus, Zika virus, Chikungunya virus, Yellow fever virus, Dengue virus, and West Nile virus, performed trainings, and deployed in Nigeria, Sierra Leone, and Senegal.
Project Sites
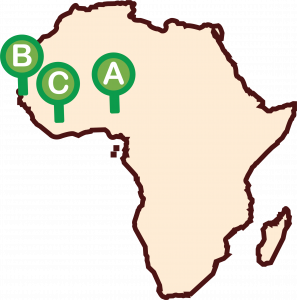
A: Nigeria
Redeemer’s University &
Irrua Specialist Teaching Hospital
B: Senegal
Université Cheikh Anta Diop
C: Sierra Leone
Kenema Government Hospital
Non-African Collaborators:
USA: Broad Institute, Harvard University, & Tulane University
Additional Resources
African Center of Excellence for Genomics of Infectious Diseases (ACEGID)






