Project Lead
The Problem
Hearing impairment (HI) is the most disabling congenital anomaly, with particularly high prevalence in sub-Saharan Africa where up to 6 / 1000 children are born deaf. It is estimated that half of children that are born with HI is because of changes in their DNA. Despite major research progresses elsewhere, genetic causes of hearing impairment in people of African ancestry are largely unknown, making this research very important. Knowing these genes can help doctors to improve their capacity to detect HI early and offer appropriate treatment and counselling to affected people and families.
Project Strategy
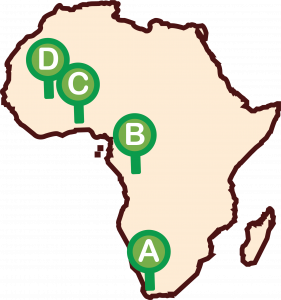
- Enroll 125 families with early-onset non syndromic hearing impairment (or NSHI) from Cameroon, Mali, Ghana, and South Africa.
- Generate next generation sequencing data on hearing-impaired family members.
- Analyze sequence data to identiy novel NSHI genes.
Potential Impact
HI-GeneS Africa will have high public health significance, especially for minority populations. Identifying novel genes can help doctors to improve their capacity to detect HI early and offer appropriate treatment and counselling to affected people and families. The project will also improve genetic screening and in the future prediction of cochlear implant and treatment outcomes in sub-Saharan Africans, African-Americans and Hispanic-Americans of African descent.